November 16th, 2010: the team composed by Andrea Pinna, Nicola Soranzo, Vincenzo De Leo and Alberto de la Fuente has qualified as the one of the best performers in the "Network inference challenge" in the DREAM5 competition which took place in the prestigious venue of Columbia University, NYC, during the annual "Joint Conference on systems biology, regulatory genomics and reverse engineering challenges". The team, called "ALF" and led by Alberto de la Fuente from the Center for Advanced Studies, Research and Development in Sardinia (CRS4), received the award also thanks to his collaboration with the Laboratory for the study of complex systems Linkalab and gained also an honorable mention.
The "Joint Conference" at Columbia University provided computational and experimental scientists, working in the field of regulatory genomics and systems biology, with the chance to compare their study approaches, aiming at a shared understanding of gene regulation mechanisms. "DREAM" stands for "Dialogue for Reverse Engineering Assessments and Methods". The goal of the challenge, at its 5th edition, is quite interesting: "Biologic network inference and quantitative modelling of biologic systems", explains Vincenzo De Leo, the researcher from Linkalab who joined ALF team, "are only at a starting point: nobody can tell how adequate their instruments are. As a consequence of that," continues De Leo, "DREAM aims at unbiased evaluation of the proposed methods".
"The teams taking part in the competition receive datasets from experiments and need to predict results that are known only by the challenge organizers", explains De Leo. "By checking how well the predictions achieved by the participants coincide with the actual results we can better understand which algorithms are more useful to obtain reliable models from genomic datasets".
Alberto de la Fuente, ALF team's scientific head, participates in DREAM competition from its second edition and already had been a best performer in DREAM challenges in 2007 and 2009. De la Fuente develops new methodologies for integrative analysis of gene expression and genotyping data to elucidate gene networks. He also focuses on the identification of dysfunctional gene networks from disease gene expression data. Vincenzo De Leo works on a project in co-operation with CRS4 and funded by Regione Sardegna, applying techniques of differential networking analysis to data about diabetes in Sardegna.
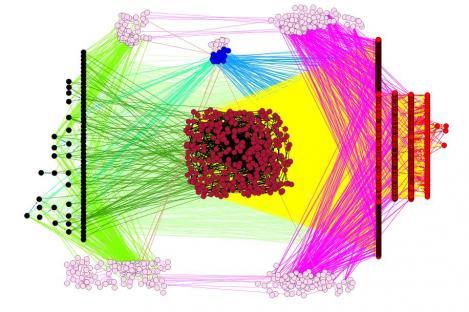
Reconstructing gene interaction complex networks is a very important step towards the comprehension of cellular behavior. The amounts of data biologists gathered in the last few years typically consist of lists of "biological objects" and information about their interactions: therefore, they are very suitable to be analyzed by Complex networks Theory.
Very often complex diseases are due to the altered interaction between sets of genes, rather than from changes occurred to a single gene. Understanding how these dynamics work using network inference can mean a progress in the research against these diseases. With the words of de la Fuente, network inference from large-scale genomics data works like that: "A cause-effect relation between the genes is established, for example applying specific gene perturbations and measuring which genes change their expression level; these genes are causally affected by the perturbed genes. Then you have to distinguish between direct and indirect cause-effect relations. The resulting network model, a Gene Network, only consists of connections with direct cause-effect relations between the genes."
29 teams from all over the world, from top universities, such as University of Chicago, Vanderbilt University, New York University and ParisTech, entered the DREAM5 network inference challenge. The challenge organizers gave the teams four datasets: three were real biologic experiments (from bacteria and yeast) and one consisted of computer-simulated experiments. Each dataset consisted of measurements obtained from several experimental situations: i.e., underexpressing or overexpressing a gene, addition of drugs or introducing changes in a time series. The aim of the research was to identify the links between transcription factors and their target genes. "The challenge organizers", explains De Leo, "gave the teams two months to solve this task: we used this time to define the techniques to obtain the best results to answer the challenge questions".
ALF team developed three different approaches to the problem; each one produced a sub-set of results. The first approach, "Perturbation response analysis", analyses of gene expression responses after removal or over-expression of single or sets of genes. The second one, "Full order partial correlation analysis of the steady-state data subset", aimed at observing the correlations among pairs of genes which could not be explained by other genes. The third approach, "Co-deviation analysis of aspecific perturbation data", aimed to identify genes which consistently deviate from their mean expression level when a given Transcription Factor deviates from its mean level. The results from the three approaches were combined to obtain a list of regulatory link predictions, which was among the best predictions in the DREAM competition.
Additional infos:
Columbia University Joint Conference website
DREAM project website
CRS4 Bioinformatics Laboratory website
Prof. Alberto de la Fuente
The poster illustrating ALF team work [file .pdf ]
Research abstract [file .pdf]
The official certificate [file .pdf]